# Basic Virology and Immunology
## **Basic Virology**
### **Introduction**
In December 2019, a series of cases of pneumonia of unknown origin were [reported](https://www.who.int/csr/don/05-january-2020-pneumonia-of-unkown-cause-china/en/) in Wuhan, the capital city of Hubei province in China. The causative virus was isolated and characterized in January 2020 ([Zhou et al., Nature 2020](https://www.nature.com/articles/s41586-020-2012-7), [Zhu et al., NEJM 2020](https://www.nejm.org/doi/10.1056/NEJMoa2001017?url_ver=Z39.88-2003\&rfr_id=ori:rid:crossref.org\&rfr_dat=cr_pub%3dwww.ncbi.nlm.nih.gov)). On January 12, 2020, the World Health Organization (WHO) tentatively named the virus as the 2019 novel coronavirus (2019-nCoV). On January 30, 2020 WHO issued a public health emergency of international concern (PHEIC) and on February 11, 2020, the WHO formally named the disease caused by the novel coronavirus as coronavirus disease 2019 (COVID-19). At that time, based on its genetic relatedness to known coronaviruses and established classification system, the International Committee on Taxonomy of Viruses [classified and renamed](https://www.nature.com/articles/s41564-020-0695-z) 2019-nCoV as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). On March 11, 2020, the [WHO formally characterized](https://www.who.int/emergencies/diseases/novel-coronavirus-2019/events-as-they-happen) the global spread of COVID-19 as a pandemic, the first to be caused by a coronavirus.
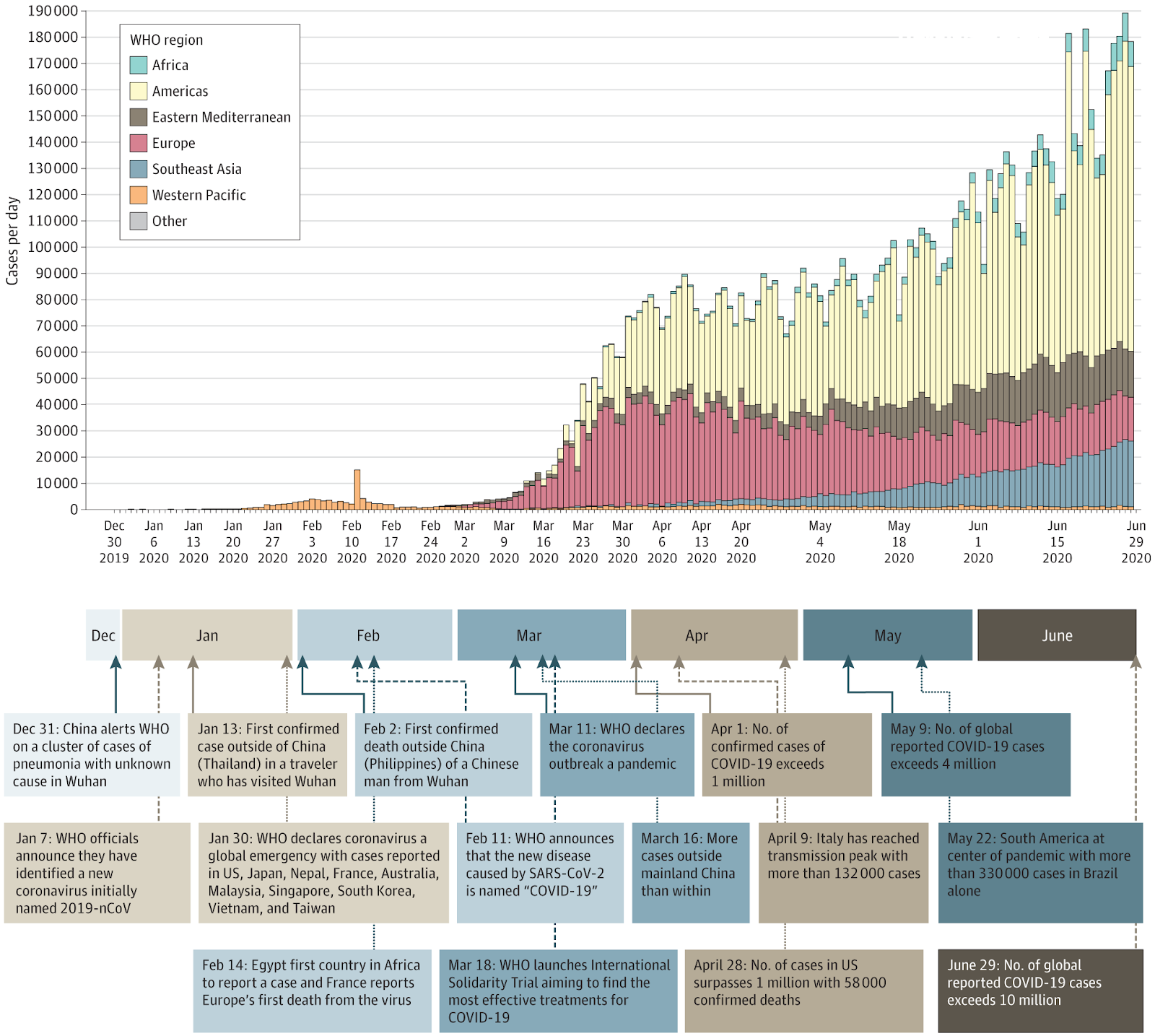
Key events in the early COVID-19 pandemic, based on data from World Health Organization situation reports. From [Wiersinga et al., JAMA, 7/10/2020](https://jamanetwork.com/journals/jama/fullarticle/2768391).
### **Classification**
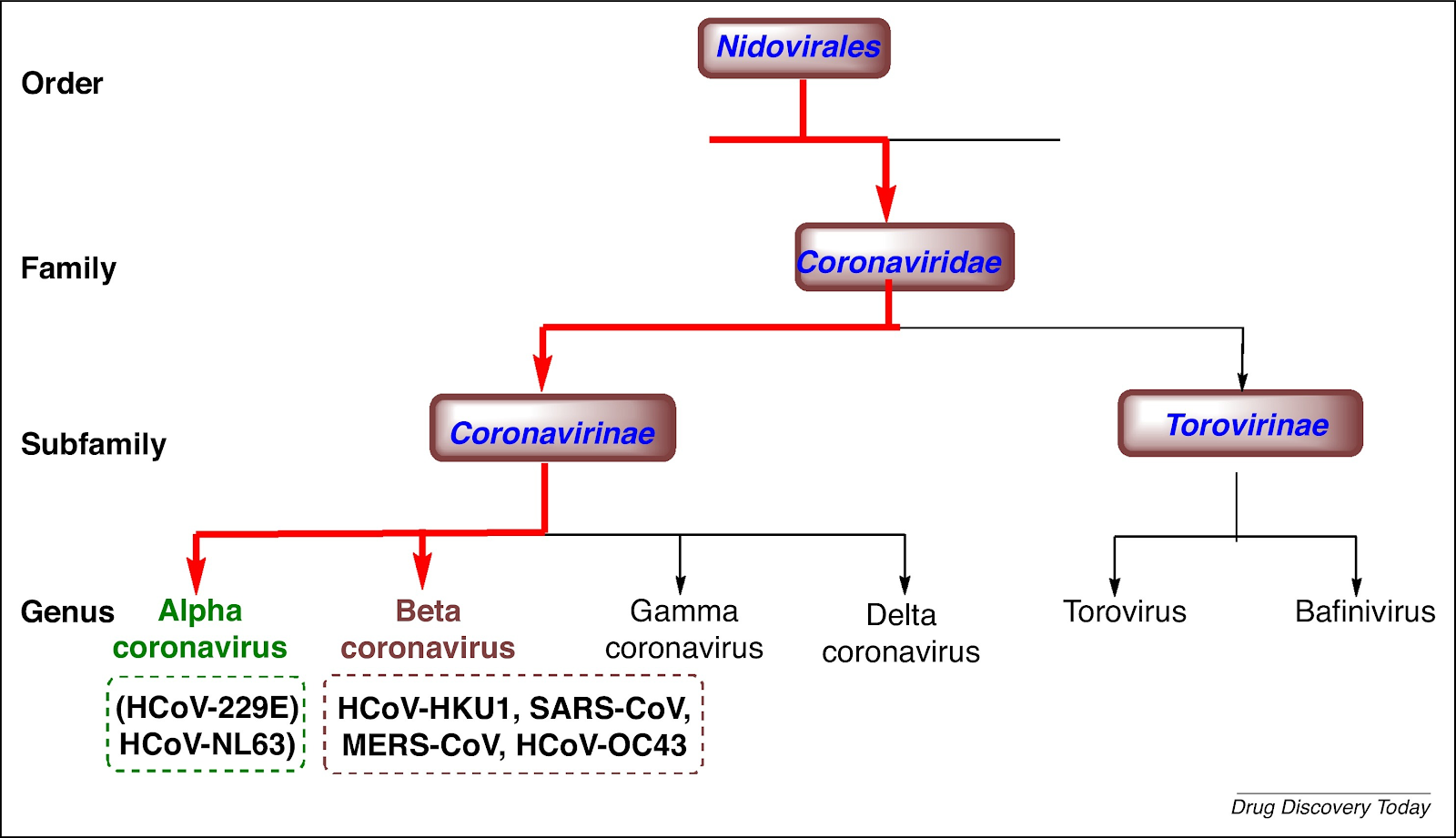
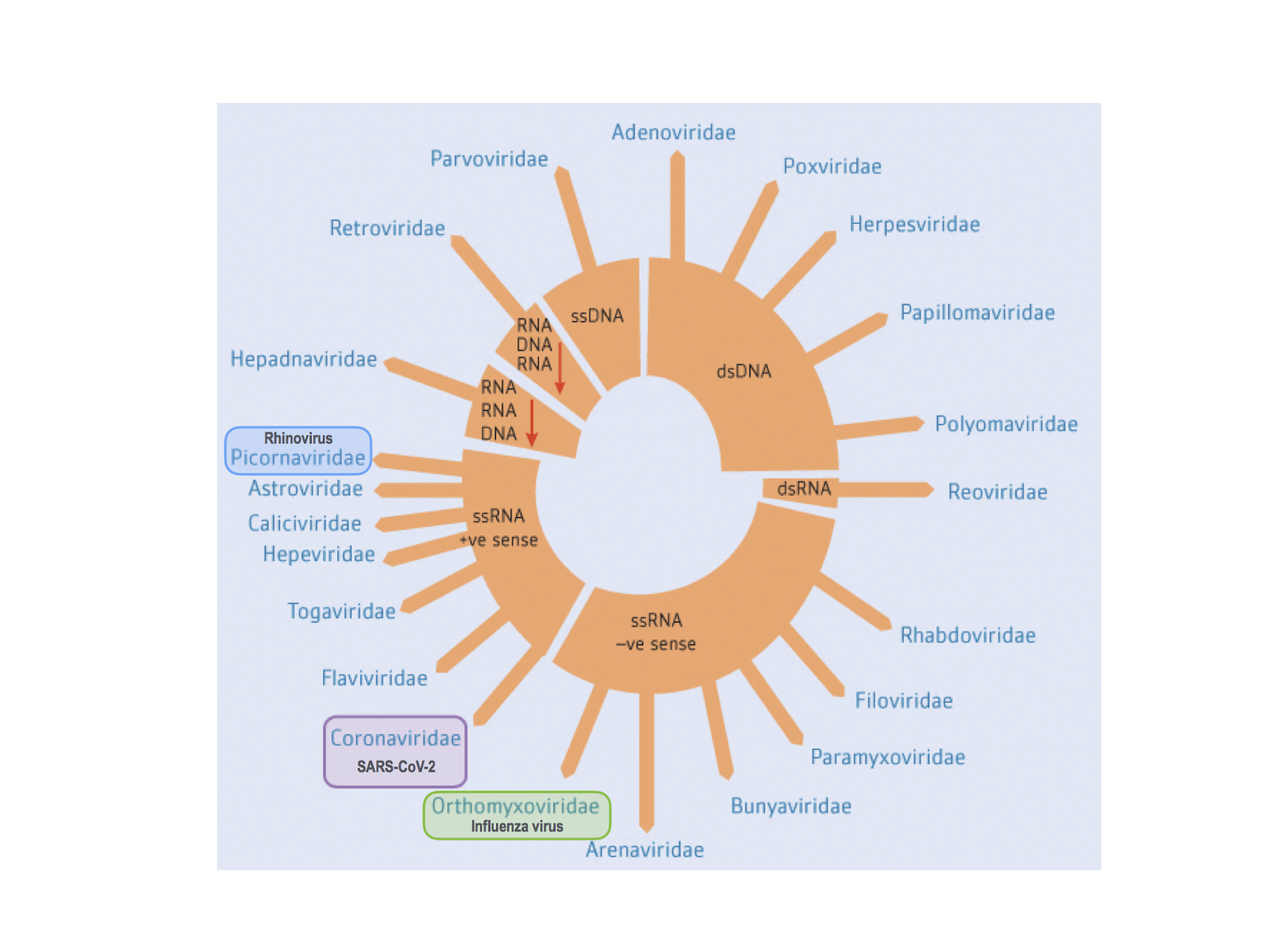
Coronaviruses are positive-sense, single-stranded enveloped RNA viruses with helical capsids that infect a wide range of hosts including humans, bats, other mammals, and birds. As shown in the schematic taxonomy below, coronaviruses are [classified](https://www.nature.com/articles/s41564-020-0695-z) within the order Nidovirales and are further subclassified into four genera: alpha, beta, delta, and gamma coronaviruses, of which alpha and beta coronaviruses are known to infect humans. As a family, coronaviruses most prominently include several human coronaviruses (HCoV) that are associated with lower pathogenicity (HCoV-229E, -NL63, -OC43, -HKU-1), contributing to seasonal cases of the ‘common cold’ and sometimes linked to more severe respiratory illness ([Bradburne et al., BMJ 1967](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1843247/?page=1); [Lieberman et al., Chest 2010](https://journal.chestnet.org/article/S0012-3692\(10\)60526-0/fulltext)). Two betacoronaviruses have previously been identified to cause more severe disease and outbreaks: severe acute respiratory syndrome coronavirus (SARS-CoV), responsible for the SARS worldwide outbreak in 2002-3 with 8,096 cases and 774 deaths [reported](https://www.who.int/csr/sars/country/table2004_04_21/en/), and Middle East respiratory syndrome coronavirus (MERS-CoV), responsible for 2,102 cases and 780 deaths [reported](https://www.who.int/emergencies/mers-cov/en/) during the 2012 MERS outbreak. SARS-CoV-2 falls within the Sarbecovirus subgenus of the Betacoronavirus genus along with SARS-CoV, and is the seventh coronavirus identified to infect humans ([Zhou et al., Nature 2020](https://www.nature.com/articles/s41586-020-2012-7), [Zhu et al., NEJM 2020](https://www.nejm.org/doi/10.1056/NEJMoa2001017?url_ver=Z39.88-2003\&rfr_id=ori:rid:crossref.org\&rfr_dat=cr_pub%3dwww.ncbi.nlm.nih.gov)).
### Genome
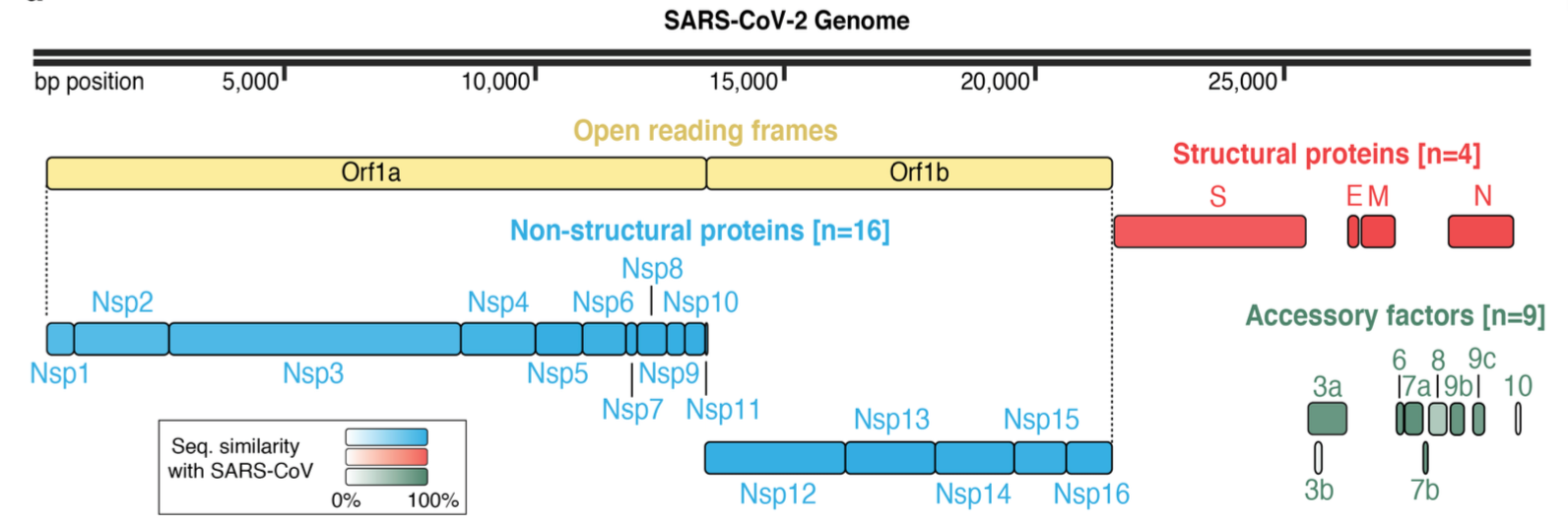
Coronaviruses have the largest genome of all ribonucleic acid (RNA) viruses infecting humans, consisting of a positive-sense single-stranded RNA roughly 30 kb in size that is 5’-capped and 3’-polyadenylated. Shown in the figure below, the virus genome is non-segmented with as many as 14 open reading frames ([Zhu et al., NEJM 2020](https://www.nejm.org/doi/10.1056/NEJMoa2001017?url_ver=Z39.88-2003\&rfr_id=ori:rid:crossref.org\&rfr_dat=cr_pub%3dwww.ncbi.nlm.nih.gov)). The genome is organized with non-structural polyproteins, which are then cleaved to be enzymes such as proteases and a RNA-dependent RNA polymerase, encoded at the 5’ end and structural proteins encoded toward the 3’ end.
### **Genetic analyses: relatedness to other viruses, suspected origins, and continued surveillance**
The majority of new coronaviruses have been isolated from bats, which serve as a natural reservoir, though other animal species have been linked as intermediate hosts in the transmission to humans, such as the palm civet and racoon dogs for SARS-CoV ([Guan et al., Science 2003](https://science.sciencemag.org/content/302/5643/276)) and dromedary camel for MERS-CoV ([Chu et al., Emerg Infect Dis 2014](https://wwwnc.cdc.gov/eid/article/20/6/14-0299_article)). Currently, the closest identified relative to SARS-CoV-2 is a virus isolated in bats with 96% sequence identity ([Zhou et al., Nature 2020](https://www.nature.com/articles/s41586-020-2012-7)). It is suspected that an intermediate host may have facilitated the zoonotic event, given both overall limited interactions between bats and humans and also the initial cluster of cases which were epidemiologically linked to a live animal and seafood market in Wuhan. Among a large variety of animals present, the pangolin, a scaly anteater and commonly trafficked mammal, has been implicated as a potential intermediate host, based on high levels of similarity of pangolin coronaviruses to SARS-CoV-2 at the protein level (Lam et al., [Nature 2020](https://www.nature.com/articles/s41586-020-2169-0)). However, full genome analysis of pangolin coronaviruses have appeared more distinct, suggesting that other market animals may have been the intermediate host between bats and humans ([Lu et al., Lancet 2020](https://www.thelancet.com/journals/lancet/article/PIIS0140-6736\(20\)30251-8/fulltext); [Zhang et al. Clin Infect Dis 2020](https://academic.oup.com/cid/advance-article/doi/10.1093/cid/ciaa112/5721420)). High levels of recombination among Sarbecoviruses and the large, unsampled diversity of coronaviruses in bats and other animals contribute to the challenge of identifying the origins of SARS-CoV-2 ([Boni et al., Nature Microbiology 2020](https://www.nature.com/articles/s41564-020-0771-4)).\
\
Continued sequencing and real-time analysis of SARS-CoV-2 genomes from samples around the world have helped track global spread, monitor local outbreaks and transmission chains, and provide insights into the epidemiology of COVID-19 ([nextstrain.org](https://nextstrain.org/narratives/ncov/sit-rep/2020-03-20?n=1)). For more information about how to read phylogenetic trees, see [here](https://nextstrain.org/narratives/trees-background/?n=1).
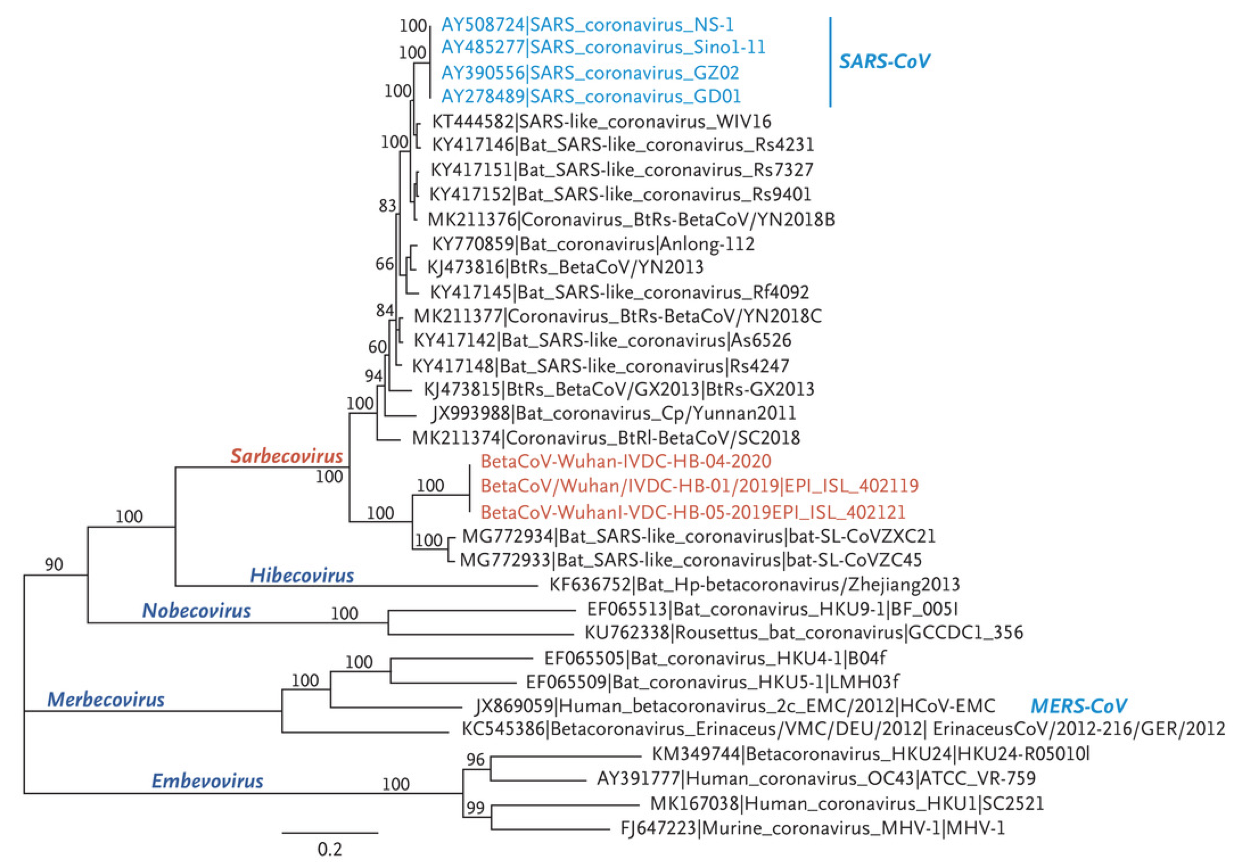
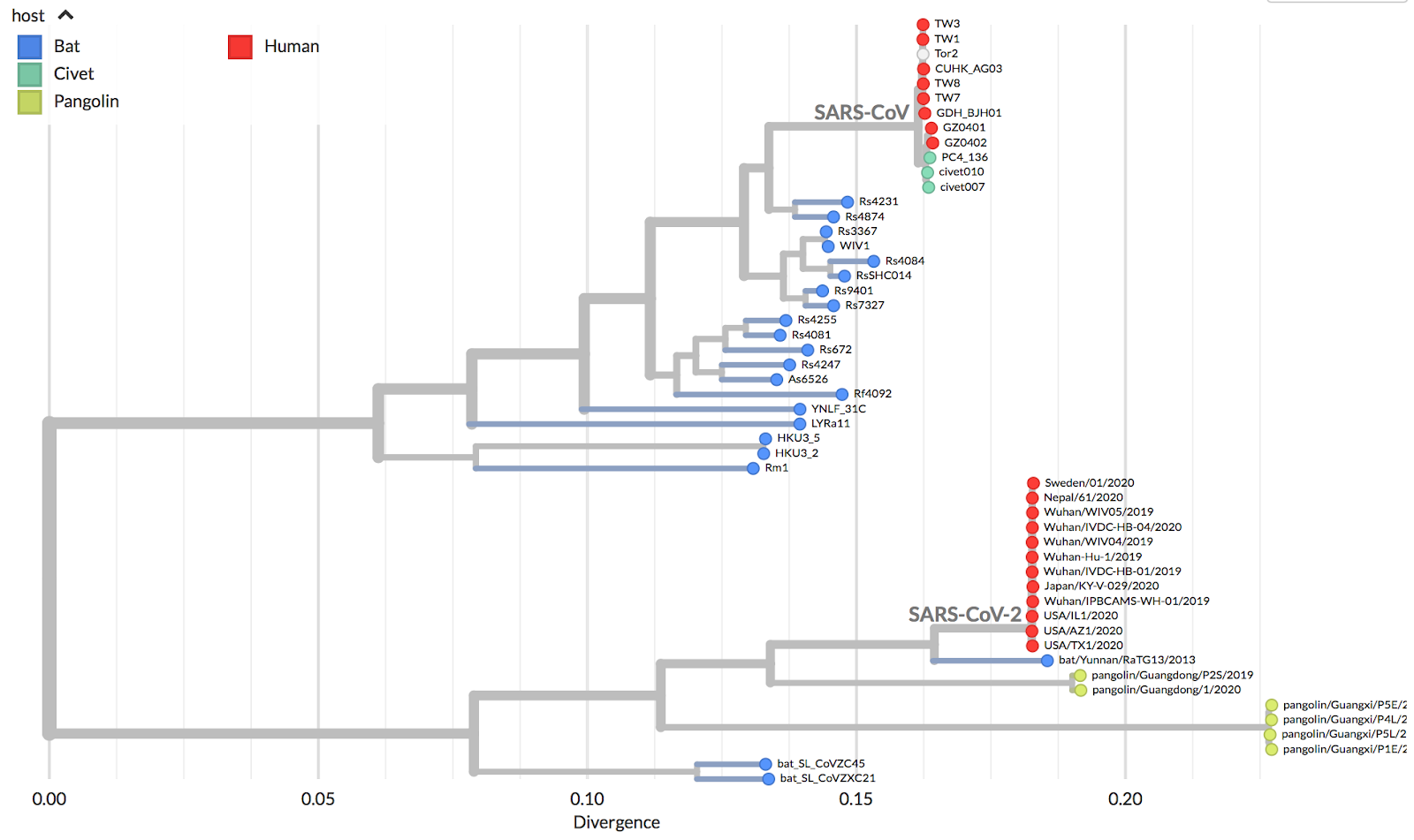
*Thought questions:*
* Based on the phylogeny, would you expect SARS-CoV-2 to behave more like SARS-CoV or MERS-CoV?
* What are some benefits and drawbacks of analyzing specific genes compared to the whole genome of a virus?
* How might understanding the origin and intermediate hosts of a virus influence human practices and policies to prevent zoonotic viruses from seeding new epidemics?
### Virus structure
Microscopically and as seen in the schematic figure below, coronaviruses have club-shaped trimeric surface spike glycoproteins that give the virions the appearance of a crown, hence their name (from the Latin corona meaning “crown”). Summarized in the table below, coronaviruses contain four major structural proteins: the spike (S), hemagglutinin-esterase (HE) in some betacoronaviruses, membrane (M), and envelope (E) all located on the membrane envelope, and the nucleocapsid (N) protein found in the core. The N proteins associate with the RNA genome to form a long helical ribonucleoprotein (RNP) packaged within the enveloped virus particle. The M protein, the most abundant of the structural proteins, is a transmembrane glycoprotein that gives the envelope its shape. The M protein embedded in the host membrane interacts with the N protein coating the viral RNA and thus helps assembly of enveloped virions. It serves as a “bridge” between the nucleocapsid and the other integral membrane proteins of the virus. The E protein is thought to be critical for coronavirus infectivity.
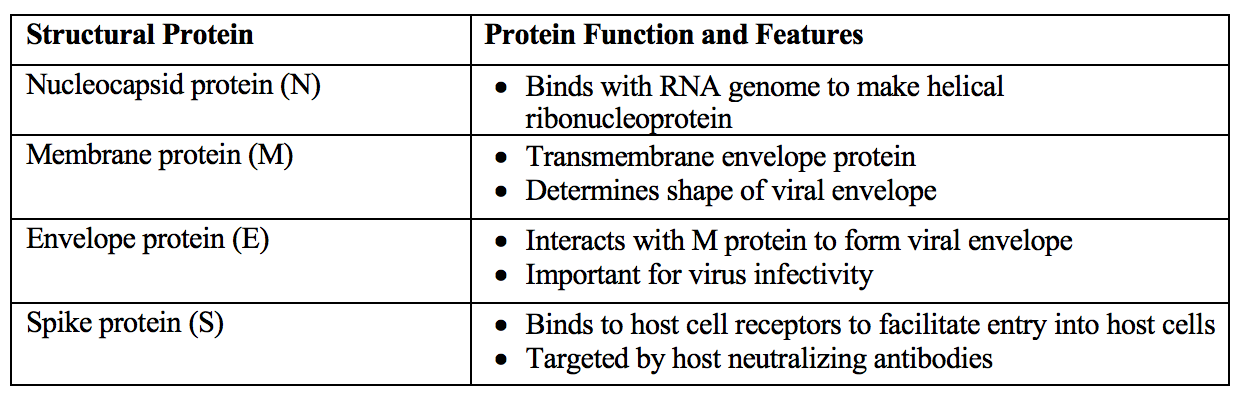
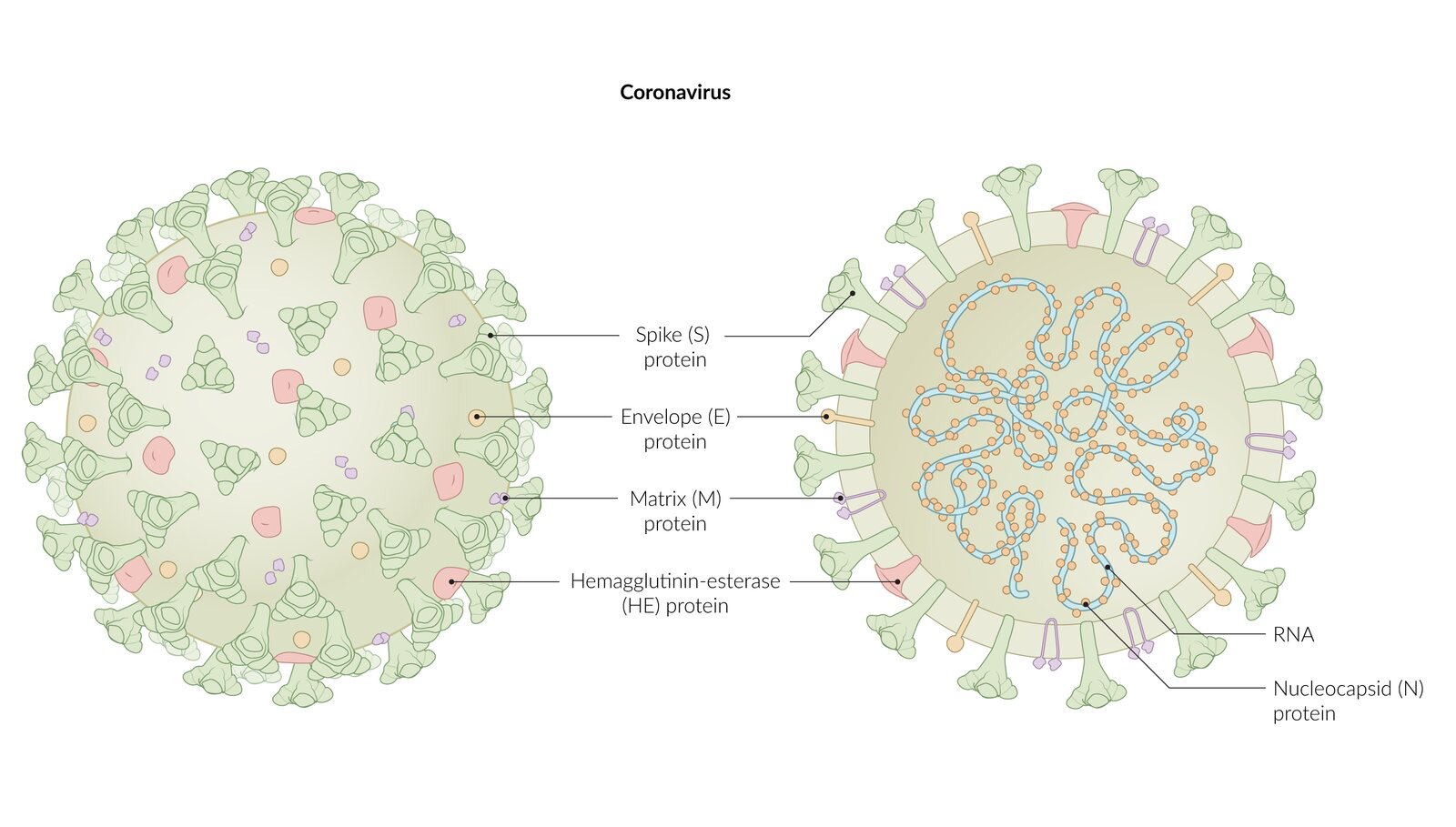
The S surface protein plays key roles in the viral life cycle and in host defense: it is responsible for receptor binding, host range, membrane fusion, hemagglutinin activity, and is a target for eliciting host neutralizing antibodies ([Millet & Whittaker, Virus Res 2015](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4465284/)). SARS-CoV-2 has genetic polymorphisms in the S protein that distinguish it from SARS- and MERS-CoV. This different spike protein structure has been suggested to allow activation by furin, a host-cell enzyme found in many human tissues including lungs, liver, and small intestines ([Andersen et al., Nature 2020](https://www.nature.com/articles/s41591-020-0820-9)). Thus, the potential for furin activation in SARS-CoV-2 may explain its expanded cellular tropism ([Walls et al., Cell 2020](https://www.cell.com/cell/fulltext/S0092-8674\(20\)30262-2?_returnURL=https%3A%2F%2Flinkinghub.elsevier.com%2Fretrieve%2Fpii%2FS0092867420302622%3Fshowall%3Dtrue)), which may contribute to the manifestation of liver injury with COVID-19 ([Zhang et al., The Lancet 2020](https://www.thelancet.com/journals/langas/article/PIIS2468-1253\(20\)30057-1/fulltext)).
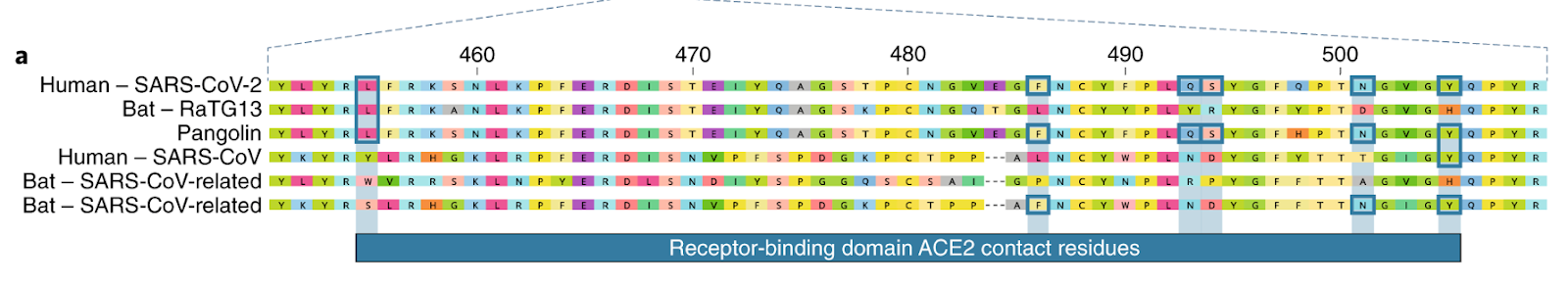
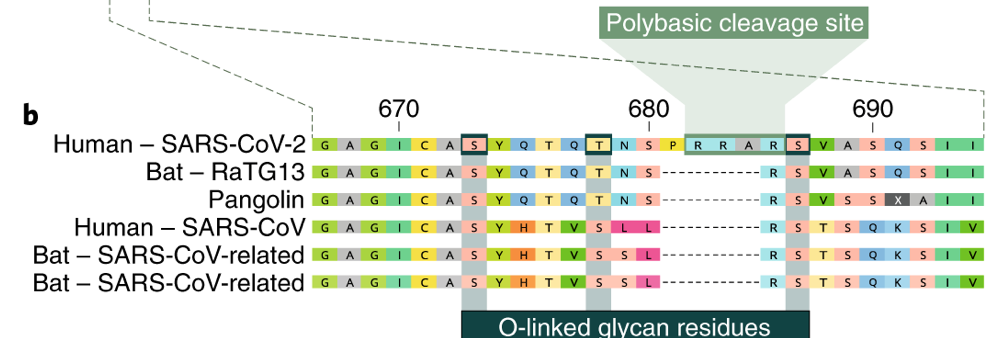
*Thought questions:*
* How could the specific proteins (S, E, M, and N) on SARS-CoV-2 be useful targets for diagnosis? For treatment? What technologies or molecular diagnostics/therapeutics would be useful?
## **Pathogenesis of COVID-19 Infection**
Research is ongoing to characterize the pathogenesis of how SARS-CoV-2 results in COVID-19 disease in humans. Below is our current understanding of the literature.
**Viral entry**
SARS-CoV-2 enters host cells through interacting with ACE2, an interferon-induced gene expressed on type 2 pneumocytes, intestinal epithelial cells, nasal goblet secretory cells ([Ziegler et al., Cell 2020](https://www.sciencedirect.com/science/article/pii/S0092867420305006)), olfactory epithelial support cells and stem cells, and nasal respiratory epithelium ([Brann et al. Science Advances 2020](https://advances.sciencemag.org/content/6/31/eabc5801)). Although SARS-CoV-2 has been found inside GI epithelial cells ([Xiao et al., Gastroenterology 2020](https://www.gastrojournal.org/article/S0016-5085\(20\)30282-1/pdf)), the virus mainly infects type 2 pneumocytes in the lung ([Zhu et al., NEJM 2020](https://www.nejm.org/doi/10.1056/NEJMoa2001017?url_ver=Z39.88-2003\&rfr_id=ori:rid:crossref.org\&rfr_dat=cr_pub%3dwww.ncbi.nlm.nih.gov)). Comprising 3% of the alveolar epithelium, type 2 pneumocytes secrete pulmonary surfactant (dipalmitoyl phosphatidylcholine), which decreases the surface tension of the lungs, and also act as stem cells for the alveolar epithelium.\
\
ACE2 is a transmembrane protein implicated in the renin-angiotensin-aldosterone system (RAAS) and hypertension pathogenesis. Note that ACE2 is a distinct enzyme from ACE: ACE converts angiotensin I to angiotensin II, a potent vasoconstrictor that drives the synthesis of aldosterone, whereas ACE2 converts active angiotensin II to angiotensin 1-7, a primary vasodilatory agent. This functions as negative regulation of RAAS. ACE2 has a protective effect in mouse models of ARDS ([Imai et al., Nature, 2005](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7094998/)). ACE2 levels in the nasal epithelium increase with age ([Bunyavanich et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2766524)), which may contribute to the differential susceptibility of older individuals to COVID-19. Of note, large studies have not shown an association with drugs that block the RAAS, such as angiotensin-converting enzyme inhibitors (ACE inhibitors) or angiotensin receptor blockers (ARBs) with increased COVID-19 acquisition or mortality ([Fosbol et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2767669)).\
\
SARS-CoV-2 binds to ACE2 via its S protein. Binding triggers a conformational change in the S protein, allowing it to be cleaved by a host cell serine protease called TMPRSS2 ([Zhou et al., Nature 2020](https://www.nature.com/articles/s41586-020-2012-7)). Cleavage of the S protein between its S1 and S2 domains allows fusion of the viral and host cell membranes and viral entry to the cell.
SARS-CoV-2 can enter by two pathways: through endocytosis, and through non-endocytic cell surface entry ([Zumla et al., Nat Rev Drug Discovery 2016](https://www.nature.com/articles/nrd.2015.37)). The endocytic pathway is a potential target of drugs like chloroquine and hydroxychloroquine ([Liu et al., Cell Discovery 2020](https://www.nature.com/articles/s41421-020-0156-0)). Upon entering in a membrane vesicle, the virion fuses with the vesicle and releases its single-segmented RNA genome into the cytosol. Since the virus is positive-sense, it can serve as mRNA and be translated immediately into non-structural viral proteins by the endogenous cell machinery. Some of these proteins form a replication complex to produce more RNA with a viral RNA-based RNA polymerase, including subgenomic RNAs which are used to translate structural proteins, and full-length transcripts to be incorporated into SARS-CoV-2 virions. The nucleocapsid (N) protein binds to the full-length positive sense viral RNA and associates with the matrix glycoprotein in the ER-Golgi intermediate compartment (ERGIC) to form a virion. The M protein associates with S and the virion virion buds into the Golgi lumen, thus gaining an envelope and exocytic vesicles from the Golgi containing the enveloped virus fuse with the cell membrane and releases viruses to infect other cells (Masters and Perlman, “Chapter 28: Coronaviridae”, Fields Virology).
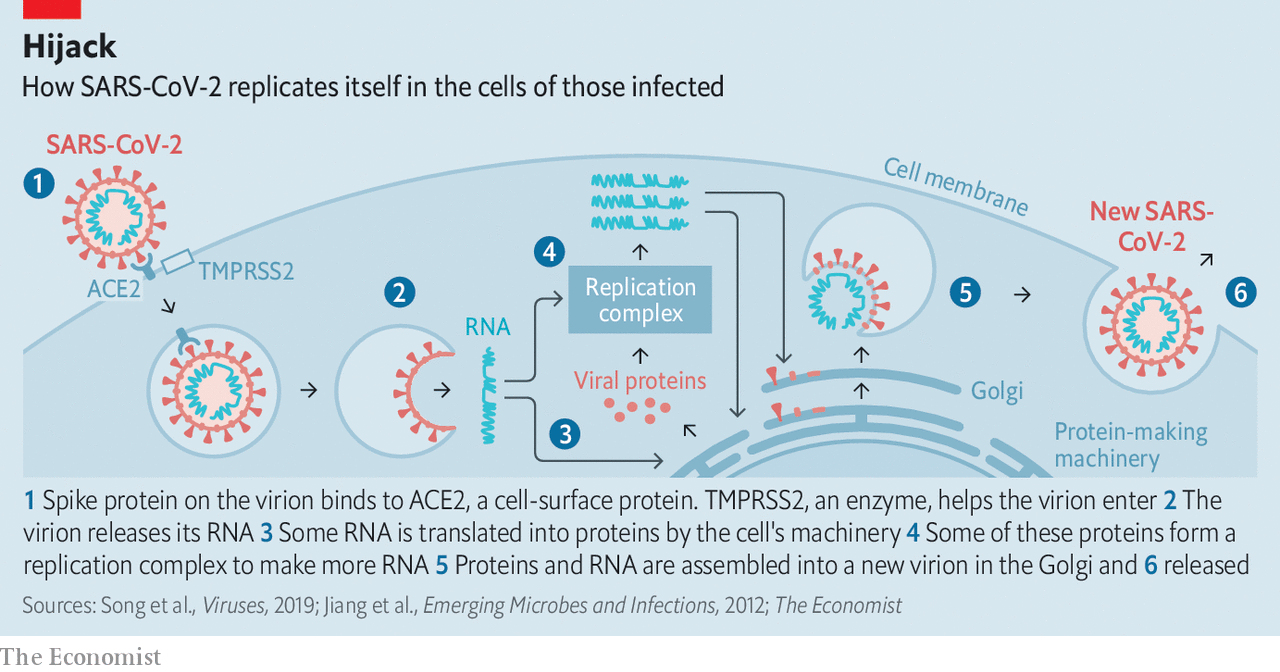
Systematic studies of possible interactions between SARS-CoV-2 and human proteins have been reported, with aims of investigating host factors mediating virus infection to identify new antiviral drug targets and repurposing of previous drugs ([Zhou et al., Cell Discovery 2020](https://www.nature.com/articles/s41421-020-0153-3); [Gordon et al., Nature 2020](https://www.nature.com/articles/s41586-020-2286-9)).
*Thought question:*
* What are the pros and cons of targeting human proteins compared to viral proteins to treat a viral disease?
**Transmission dynamics**
Transmission of SARS-CoV-2 is thought to occur mainly through respiratory droplets ([Aylward et al., Report of the WHO-China Joint Mission 2020](https://www.who.int/docs/default-source/coronaviruse/who-china-joint-mission-on-covid-19-final-report.pdf)). Prolonged exposures to an asymptomatic infected person (within 6 feet for at least 15 minutes) and briefer exposures to symptomatic individuals (coughing) are associated with higher risk of transmission than shorter exposures to asymptomatic individuals ([Chu et al., Lancet 2020](https://www.thelancet.com/pdfs/journals/lancet/PIIS0140-6736\(20\)31142-9.pdf)). Other routes of transmission such as virus contamination of common objects and aerosolization in a confined space have been suggested, though the significance of their role in contributing to overall transmission have yet to be fully elucidated ([Cai et al., Emerg Inf Dis 2020](https://wwwnc.cdc.gov/eid/article/26/6/20-0412_article); [Rothe et al., NEJM 2020](https://www.nejm.org/doi/full/10.1056/NEJMc2001468)). Respiratory droplets can be generated by sneezing (40,000 droplets), coughing (3,000 droplets), or talking (about 600 droplets per minute). They can also be produced by medical procedures like intubation and bronchoscopy or by use of oxygen masks and nebulizers ([Tang et al., Journal of Hospital Science 2006](https://www.sciencedirect.com/science/article/pii/S0195670106002866)). Larger droplets (>5 microns) tend to spread about 1-2 meters (3-6 feet) before the force of gravity causes them to drop to the ground and require droplet precautions. Aerosols are smaller and rapidly evaporate, leaving behind droplet nuclei that can spread further and remain suspended in the air for hours; these require stricter airborne precautions ([Klompas et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2768396)). While experimental data has shown that SARS-CoV-2 can remain suspended in the air and viable for many hours and that aerosolization can occur outside of aerosolizing procedures, epidemiological data and secondary attack rates support a primarily droplet based mode of transmission, although rare events of transmission via airborne routes have been documented in settings like choir practice and a call center ([Klompas et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2768396)). To reduce the risk of exposure, current guidelines recommend physical distancing and mask wearing. A systematic review and meta-analysis of studies on SARS, MERS and COVID-19 found that physical distancing of more than 3 feet from others had an adjusted odds ratio of coronavirus transmission of 0.18 ([Chu et al., Lancet 2020](https://www.thelancet.com/pdfs/journals/lancet/PIIS0140-6736\(20\)31142-9.pdf)). The CDC currently recommends staying at least 6 feet away from others who do not live in the same household (). This topic remains an area of active investigation.
Reports indicate that SARS-CoV-2 has the potential to be transmitted through [fomites](https://en.wikipedia.org/wiki/Fomite), or objects with virus on their surface, although this route of transmission is less important compared to respiratory transmission. SARS-CoV-2 appears to have similar viability in aerosols and on surfaces when compared to that of SARS-CoV. When aerosolized, SARS-CoV-2 remains viable for up to 3 hours, a critical consideration for hospital infection control, particularly when undergoing aerosolizing procedures. Viable SARS-CoV-2 was measured from surfaces up to 4 hours on copper, 24 hours on cardboard, and 72 hours on plastic and stainless steel ([van Doremalen et al., NEJM 2020](https://www.nejm.org/doi/full/10.1056/NEJMc2004973)). While these results do not fully evaluate the infectivity of the virus on different surfaces, in general this data supports the notion that maintaining good hygiene (washing hands often, especially after touching public surfaces, and avoiding touching face and mouth) could help mitigate the spread of SARS-CoV-2. Moreover, this data suggests that other viral properties must explain the infectivity differences between SARS-CoV-2 and SARS-CoV. Previous work studying other coronaviruses have also suggested that surface disinfection such as with 62-71% ethanol or 0.5% hydrogen peroxide, commonly found in household cleaning products, can inactivate coronaviruses that persist on surfaces ([Kampf et al., J Hosp Infect 2020](https://www.journalofhospitalinfection.com/article/S0195-6701\(20\)30046-3/fulltext)).
Mask wearing is important to lessen transmission from asymptomatic, presymptomatic and symptomatic individuals alike, and helps protect both the mask wearer and those around them. A meta-analysis of studies on SARS, MERS and COVID-19 found that mask wearing resulted in an adjusted odds ratio of 0.15 (0.07-0.34) of infection (85% decreased risk with face masks compared to no face masks); N95 masks were associated with an adjusted odds ratio of 0.04, while surgical and cloth face masks had an adjusted odds ratio of 0.33 ([Chu et al., Lancet 2020](https://www.thelancet.com/pdfs/journals/lancet/PIIS0140-6736\(20\)31142-9.pdf)). In the same study, eye protection was associated with an adjusted odds ratio of 0.22 compared to no eye protection. Surgical masks have shown efficacy in hospitalized patients with seasonal coronaviruses ([Leung et al., Nat Med 2020](https://www.nature.com/articles/s41591-020-0843-2)), and homemade cloth masks can block large droplets produced during speech ([Anfinrud et al., MedRXiv 2020](https://www.medrxiv.org/content/10.1101/2020.04.02.20051177v1)). Current CDC recommendations are for the lay public to wear cloth masks in public settings and when around other people who don’t live in the same household and to reserve N95 masks for people with high levels of exposure ([cdc.gov](https://www.cdc.gov/coronavirus/2019-ncov/prevent-getting-sick/cloth-face-cover.html)).
In concordance with the ability of SARS-CoV-2 to infect intestinal epithelial cells, viral RNA has been detected in 29–55% of stool samples from COVID-19 patients ([Xiao et al., Gastroenterology 2020](https://www.gastrojournal.org/article/S0016-5085\(20\)30282-1/pdf);[ Wang et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2762997); [Wu et al., Lancet Gastroenterol Hepatol 2020](https://www.thelancet.com/journals/langas/article/PIIS2468-1253\(20\)30083-2/fulltext)), as well as environmental samples from the toilet ([Ong et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2762692)). Live virus has also been isolated from stool specimens ([Wang et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2762997); [Zhang et al., CCDC Weekly 2020](http://weekly.chinacdc.cn/en/article/id/ffa97a96-db2a-4715-9dfb-ef662660e89d)). Viral RNA can be detected in the stool or on rectal swabs even after oro-/nasopharyngeal swabs turn negative ([Xiao et al., Gastroenterology 2020](https://www.gastrojournal.org/article/S0016-5085\(20\)30282-1/pdf); [Xu et al., Nat Med 2020](https://www.nature.com/articles/s41591-020-0817-4)). These data raise the possibility of fecal–oral transmission of SARS-CoV-2 ([Yeo et al., Lancet Gastroenterol Hepatol 2020](https://www.thelancet.com/journals/langas/article/PIIS2468-1253\(20\)30048-0/fulltext)), as was suspected with the SARS outbreak of 2002–2003 ([Abdullah et al., Emerg Inf Dis 2003](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3016765/)). Live SARS-CoV-2 has also been isolated from blood but only from rare patients, and RNA has been isolated from the conjunctiva of the eyes and from urine samples ([Wang et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2762997);[ Liang & Wu, Acta Opthalm 2020](https://onlinelibrary.wiley.com/doi/full/10.1111/aos.14413); [Peng et al., J Med Virol 2020](https://pubmed.ncbi.nlm.nih.gov/32330305/)).
COVID-19 can be spread by vertical transmission, passed from mother to fetus or neonate during pregnancy or during the perinatal period, but this occurrence is very rare. A study of 31 deliveries in women with SARS-CoV-2 infection found evidence of infection in two newborn infants, with only one episode of confirmed congenital infection prior to delivery ([Fenizia et al., Nat Comm 2020](https://www.nature.com/articles/s41467-020-18933-4)). In a number of limited case series of pregnant women with lab-confirmed COVID-19, none of the infants were found to have COVID-19, and SARS-CoV-2 was not detected in samples including amniotic fluid, cord blood, neonatal throat swab, or breastmilk ([Chen et al., Lancet 2020](https://www.thelancet.com/journals/lancet/article/PIIS0140-6736\(20\)30360-3/fulltext#seccestitle130), [Li et al., Emerg Infect Dis 2020](https://wwwnc.cdc.gov/eid/article/26/6/20-0287_article), [Schwartz, Arch Path Lab Med 2020](https://www.archivesofpathology.org/doi/10.5858/arpa.2020-0901-SA)). Previous limited case series have found infants born to mothers with SARS were negative for SARS-CoV ([Wong et al., Am J Ob Gyn 2004](https://www-sciencedirect-com.ezp-prod1.hul.harvard.edu/science/article/pii/S0002937803020398); [Shek et al., Pediatrics 2003](https://pediatrics.aappublications.org/content/pediatrics/112/4/e254.full.pdf?ck=nck)) and vertical transmission with SARS or MERS infection have not been documented in the past ([Schwartz & Graham, Viruses 2020](https://www.mdpi.com/1999-4915/12/2/194/htm)). Cases of COVID-19 have been reported in neonates and infants; however, these were complicated by close contact history with confirmed infected persons following birth ([Qiao, Lancet 2020](https://www.thelancet.com/journals/lancet/article/PIIS0140-6736\(20\)30365-2/fulltext); [Wei et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2761659)). In contrast, neonates born to COVID-positive mothers can acquire IgG (transplacentally) and IgM (mechanism unknown, possibly imperfect assay) antibodies against the disease ([Zeng et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2763854)).
A feature of COVID-19 is its ability to be transmitted by asymptomatic individuals, whether before symptoms start or by individuals who do not have symptoms. Viral shedding appears to start 2 to 3 days prior to the onset of symptoms and peaks around the time of symptom onset ([Aylward et al., Report of the WHO-China Joint Mission 2020](https://www.who.int/docs/default-source/coronaviruse/who-china-joint-mission-on-covid-19-final-report.pdf); [He et al., Nature Medicine 2020](https://www.nature.com/articles/s41591-020-0869-5)), and has been shown in asymptomatic individuals ([Zhou et al., NEJM 2020](https://www.nejm.org/doi/full/10.1056/NEJMc2001737)). Viral load can be positive multiple (1 to 7) days before symptom onset ([Wang et al., JID 2020](https://academic.oup.com/jid/advance-article/doi/10.1093/infdis/jiaa119/5807958)), and peaks around the time of symptom onset, suggesting significant viral load before someone knows they may be infected ([To et al., Lancet ID 2020](https://www.thelancet.com/journals/laninf/article/PIIS1473-3099\(20\)30196-1/fulltext)). Modeling of transmission events in China prior to the January 23rd travel restrictions estimated that undocumented cases, which experienced no to mild symptoms that did not warrant hospitalization, were responsible for 79% of new cases ([Li et al., Science 2020](https://science.sciencemag.org/content/early/2020/03/13/science.abb3221)); other studies found that 12-62% of cases were transmitted from pre-symptomatic individuals ([Du et al., Emerging Inf Dis, 2020](https://wwwnc.cdc.gov/eid/article/26/6/20-0357_article); [Ganyani et al., Eurosurveillance 2020](https://www.eurosurveillance.org/content/10.2807/1560-7917.ES.2020.25.17.2000257)). Current estimates of the COVID-19 incubation period, which refers to the time period from initial exposure to symptom onset, range from 1-14 days with a median of 5 days and 95th percentile of 12 days, similar to SARS ([Lauer et al., Ann Intern Med 2020](https://annals.org/aim/fullarticle/2762808/incubation-period-coronavirus-disease-2019-covid-19-from-publicly-reported); [Li et al., NEJM 2020](https://www.nejm.org/doi/10.1056/NEJMoa2001316?url_ver=Z39.88-2003\&rfr_id=ori:rid:crossref.org\&rfr_dat=cr_pub%3dwww.ncbi.nlm.nih.gov)). The detection of SARS-CoV-2 RNA in patients at 20 days and as long as 37 days also suggests the potential of prolonged virus shedding ([Zhou et al., Lancet 2020](https://www.thelancet.com/journals/lancet/article/PIIS0140-6736\(20\)30566-3/fulltext#seccestitle10); [He et al., Nature Medicine 2020](https://www.nature.com/articles/s41591-020-0869-5)). Increased severity of cases have been suggested to be associated with higher viral loads and longer duration of viral shedding ([Liu et al., Lancet Inf Dis 2020](https://www.thelancet.com/journals/laninf/article/PIIS1473-3099\(20\)30232-2/fulltext)). Because they may shed virus for a period of time after symptoms have resolved, it is still unknown how long someone in remission from COVID-19 remains infectious. However, for people who become symptomatic, a recent study was not able to isolate infectious virus after 8 days after symptom onset, suggesting a limited time period for infectious viral shedding ([Bullard et al., Clin Inf Dis 2020](https://academic.oup.com/cid/advance-article/doi/10.1093/cid/ciaa638/5842165)). Truly asymptomatic cases are likely relatively rare, with rates of asymptomatic cases estimated to make up 15% of total cases overall, with more asymptomatic younger individuals than older ones ([Byambasuren et al., JAMMI Canada 2020](https://jammi.utpjournals.press/doi/10.3138/jammi-2020-0030)). The delay between exposure and showing symptoms combined with transmission from asymptomatic or presymptomatic hosts have made SARS-CoV-2 particularly difficult to contain.
**COVID-19 Variants**
Like all viruses, COVID-19 changes through mutation as it replicates through populations. Multiple COVID-19 variants have emerged, some more consequential than others. Notable variants include a new variant called B.1.1.7 that emerged in the U.K. and has been identified in numerous countries including the U.S. The mutation affects an amino acid in the receptor binding domain of the spike protein, the portion that physically interacts with ACE2. This variant is known to spread more easily than previous variants leading to projections that it will be the dominant strain in the US by March. The CDC has emphasized that while there is no evidence that this strain is more deadly than previous strains, increased transmission will equal more cases and subsequently more deaths ([MMWR CDC 1.15.21](https://www.cdc.gov/mmwr/volumes/70/wr/mm7003e2.htm?s_cid=mm7003e2_w)). However the current vaccines will likely still be protective against this variant. Additional variants that have been identified in the US starting at the end of January 2021 include the 1.351 variant in South Africa and the P.1 variant in Brazil.\
\
Further discussion of COVID-19 variants to watch can be found on the regularly updated [CDC’s New Variants page](https://www.cdc.gov/coronavirus/2019-ncov/transmission/variant.html).
*Thought questions:*
* How would you predict that a difference in infected cell types might change the presentation and transmission of COVID-19?
* Imagine a few real-life scenarios that you may soon encounter or may have already encountered:
* [**Diane** ](https://curriculum.covidstudentresponse.org/curriculum-overview/cases)wants to order food from a delivery service, but is worried about getting sick. What advice would you give her about touching packages, meeting the delivery person, and ordering premade food?
* [**Brian**](https://curriculum.covidstudentresponse.org/curriculum-overview/cases) has a friend who had low fevers, fatigue, and a dry cough, but was never tested for COVID-19. His friend self-quarantined at home and now has not had any symptoms for the past day. Brian wants to hang out with this friend today. What would you tell him about his risk of exposure? What precautions should he take if he decides to see his friend?
* How do estimates of incubation periods and viral shedding inform public health efforts?
**Pathogenesis of Organ Damage in COVID-19**
Like pulmonary epithelial cells, vascular endothelial cells express ACE2, and SARS-COV-2 has been found inside of endothelial cells in pulmonary capillaries, leading to inflammatory cytokine production, endothelial cell death, and endothelial barrier disruption ([Varga et al., Lancet 2020](https://www.thelancet.com/pdfs/journals/lancet/PIIS0140-6736\(20\)30937-5.pdf)). This endotheliitis is associated with diffuse thickening of the alveolar wall and infiltration of mononuclear cells and macrophages into the airspaces, which collectively are seen as ground-glass opacities in the lungs on CT imaging ([Wiersinga et al., JAMA, 7/10/2020](https://jamanetwork.com/journals/jama/fullarticle/2768391)). As the inflammation progresses, pulmonary edema and hyaline membrane formation occur and cause acute respiratory distress syndrome (ARDS), which interferes with oxygen diffusion.
Activation of the coagulation cascade and consumption of clotting factors have been associated with severe COVID-19. In one study, 71% of patients who died of COVID-19 met criteria for disseminated intravascular coagulation (DIC) ([Tang et al., J Thrombosis Haemostasis 2020](https://onlinelibrary.wiley.com/doi/full/10.1111/jth.14768)); in another, 31% of COVID-19 patients admitted to ICUs had at least one thrombotic complication ([Klok et al., Thrombosis Res 2020](https://www.thrombosisresearch.com/article/S0049-3848\(20\)30120-1/pdf)). SARS-CoV-2 does not appear to have intrinsic procoagulant activity; instead, the increased coagulation is likely due to a combination of endothelial damage and procoagulopathic effects of the inflammatory response itself ([Connors and Levy, Blood 2020](https://ashpublications.org/blood/article/135/23/2033/454646/COVID-19-and-its-implications-for-thrombosis-and)).
Neurological symptoms such as myalgias, headaches, encephalopathy, dizziness, dysgeusia (loss of taste), and anosmia (loss of smell) are common in COVID-19 patients, affecting up to 82.3% of COVID-19 patients at some point in their disease course in one study ([Liotta et al., Annals of Clinical and Translational Neurology 2020](https://onlinelibrary.wiley.com/doi/full/10.1002/acn3.51210)). So far, there are several theories proposed to explain the cause of COVID-associated neurological symptoms. The loss of smell and taste may be explained by infection of non-neuronal ACE2-positive cells in the nasal and oral mucosa, such as endothelial cells, oligodendrocytes, and astrocytes. This can alter axon conduction velocity, metabolic and neurotransmitter homeostasis, and/or cerebral perfusion ([Pan et al., Academic Radiology 2020](https://www.academicradiology.org/article/S1076-6332\(20\)30507-9/fulltext)). The SARS-CoV-2 mediated cytokine storm can injure the blood-brain barrier, which allows the entry of cytokines into the brain, resulting in seizures and encephalopathy. SARS-CoV-2 activated cytokines can also cause immune-mediated “molecular mimicry” which damages cranial nerves, peripheral nerves, and/or muscles ([Majid et al., Journal of Alzheimer’s Disease 2020](https://content.iospress.com/articles/journal-of-alzheimers-disease/jad200581)).
Lingering symptoms are common in people who have had COVID-19, even in those who had mild or asymptomatic disease. These symptoms have been described as “long COVID”, “post-COVID syndrome”, or “post-acute COVID-19 syndrome” ([IDSA 11/19/20](https://www.idsociety.org/covid-19-real-time-learning-network/disease-manifestations--complications/post-covid-syndrome/)); the lay press has called people with these symptoms “long haulers” ([Yong, Atlantic 8/19/20](https://www.theatlantic.com/health/archive/2020/08/long-haulers-covid-19-recognition-support-groups-symptoms/615382/)). Studies following up on patients weeks to months after diagnosis or hospitalization with COVID-19 have found that 35%-87% of patients had ongoing symptoms, with fatigue, dyspnea, anosmia, dysgeusia, and weakness frequently reported ([Tenforde et al., MMWR 2020](https://www.cdc.gov/mmwr/volumes/69/wr/mm6930e1.htm); [Halpin, J Med Virol 2020](https://onlinelibrary.wiley.com/doi/full/10.1002/jmv.26368); [Carfi et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2768351); [Garrigues, J Infection 2020](https://www.journalofinfection.com/article/S0163-4453\(20\)30562-4/fulltext); [Carvalho-Schneider, Clin Micro Infection 2020)](https://www.clinicalmicrobiologyandinfection.com/article/S1198-743X\(20\)30606-6/fulltext). In one study, 66% of adults with COVID-19 who had not been admitted to an ICU reported persistent symptoms at day 60 after first positive RT-PCR test ([Carvalho-Schneider, Clin Micro Infection 2020](https://www.clinicalmicrobiologyandinfection.com/article/S1198-743X\(20\)30606-6/fulltext)). A recent study found a high prevalence of autoantibodies in people with both mild/moderate and severe COVID-19, including some that target immunological functions and can worsen severity of disease in a mouse model of COVID-19; other autoantibodies targeted proteins found in the central nervous system, vascular cells, and connective tissue ([Wang, MedRXiv 12/12/20](https://www.medrxiv.org/content/10.1101/2020.12.10.20247205v3)). These autoantibodies may provide a link between COVID-19 infection and the autoimmune-like symptoms found in some individuals.
## Immune Response in COVID-19
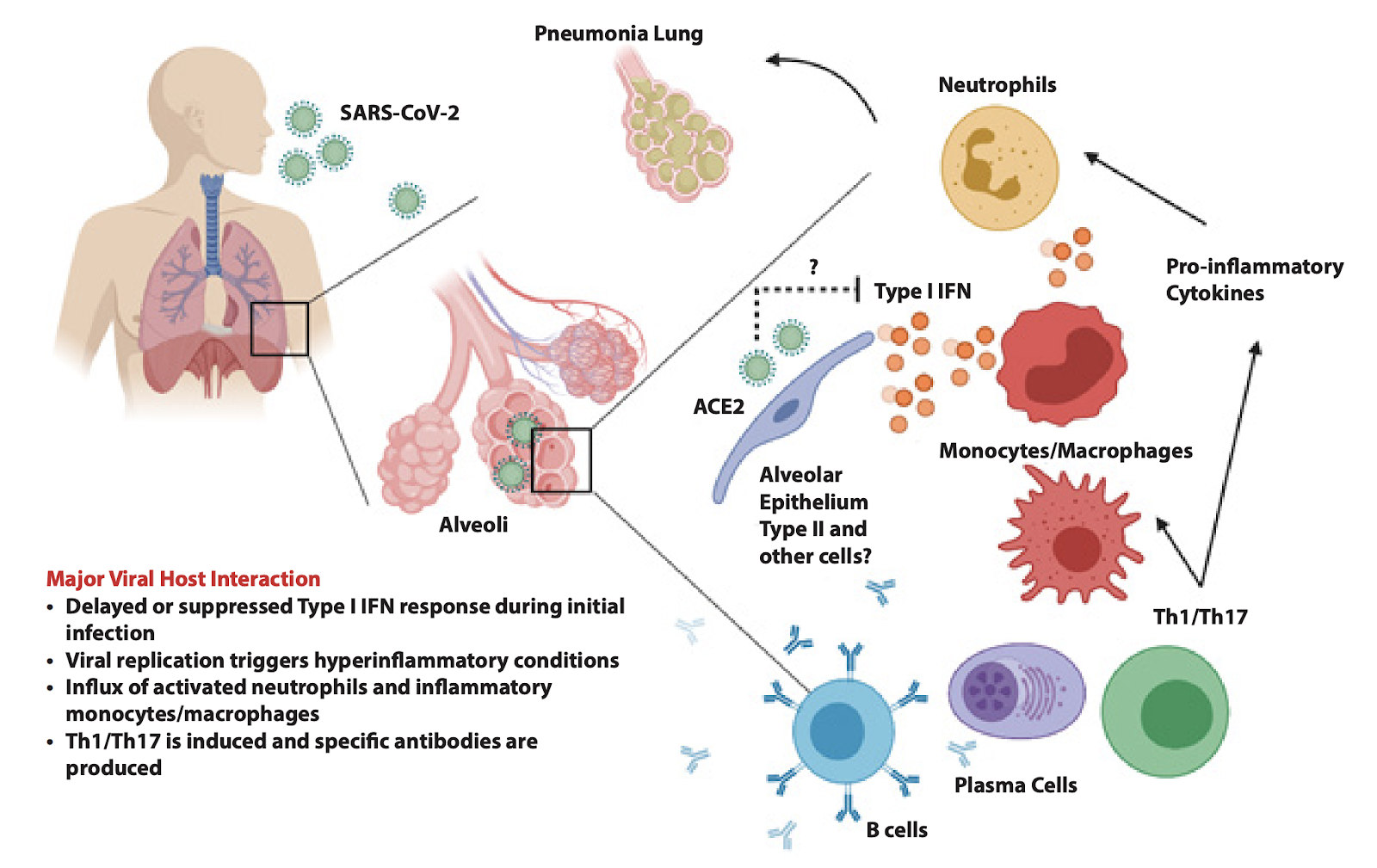
### Innate immune response
In severe cases of COVID-19, decreased viral control of SARS-CoV-2 is associated with a delayed or absent type I interferon response ([Chu et al., Clin Inf Dis, 2020](https://academic.oup.com/cid/advance-article/doi/10.1093/cid/ciaa410/5818134)); instead, the initial response recruits neutrophils, monocytes and macrophages to the lung, which is associated with increased immunopathology ([Blanco-Melo et al., Cell 2020](https://www.cell.com/pb-assets/products/coronavirus/CELL_CELL-D-20-00985.pdf); [Hadjadj et al., MedRXiv preprint 2020](https://www.medrxiv.org/content/10.1101/2020.04.19.20068015v1)). Like many other RNA viruses, SARS-CoV-2 RNA is detected by cytosolic sensors including RIG-I and MDA-5, which then interact with MAVS on the mitochondrial surface; downstream signaling leads to transcription of type I interferon genes. IFN-I binds to the interferon alpha receptor (IFNAR), which leads to expression of interferon stimulated response genes that create an antiviral state. At least 10 SARS-CoV-2 viral proteins inhibit elements of the cellular type I interferon response, including modifying viral RNA to make it difficult to detect, leading to degradation of RIG-I and MAVS, blocking host translation, and promoting IFNAR1 degradation ([Sa Ribero et al., PLoS Pathogens 2020](https://journals.plos.org/plospathogens/article?id=10.1371/journal.ppat.1008737)). SARS-CoV-2 induces a less robust interferon response than SARS-CoV, which may explain the delay in symptom onset in COVID-19 compared to SARS. Additionally, two recent studies ([Bastard et al., Science 2020, Zhang et al., Science 2020](https://science.sciencemag.org/content/early/2020/09/23/science.abd4585)) show that some patients harbor neutralizing auto-antibodies against type I interferons or mutations in genes associated with type I interferon signaling, and are both overrepresented in about 14% of people who developed severe COVID-19 disease.
The influx of myeloid cells into the lungs is accompanied by increases in levels of serum pro-inflammatory cytokines, such as IL-1, IL-6, IL-12 and TNFɑ, that increase vascular permeability and decrease lung function. IL-6 can signal through direct binding to the IL-6 receptor on lymphocytes; it can also bind to soluble IL-6 receptor and bind to endothelial cells to stimulate vascular effects of the disease, and finally it can change expression of inflammatory mediators in the liver ([Moore and June, Science 2020](https://science.sciencemag.org/content/368/6490/473/tab-pdf)). In COVID-19, both increases in monocytes and neutrophils in the lungs ([Prompetchara et al., Asian Pacific Journal of Allergy and Immunology 2020](http://apjai-journal.org/wp-content/uploads/2020/02/Covid_AP-200220-0772.pdf)) and higher serum pro-inflammatory responses are associated with severe disease ([Huang et al., Lancet 2020](https://www.thelancet.com/journals/lancet/article/PIIS0140-6736\(20\)30183-5/fulltext)). However, when levels of these cytokines were compared between critically ill patients with COVID-19 and other conditions like septic shock with or without ARDS, the COVID patients had lower levels of TNFa, IL-6 and IL-8 than septic patients with ARDS and lower levels of IL-6 and IL-8 than septic patients without ARDS ([Kox et al., JAMA 2020](https://jamanetwork.com/journals/jama/fullarticle/2770484)), which calls into question the extent of a “cytokine storm” in COVID-19 compared to other critical illnesses. It is thought that glucocorticoids like dexamethasone, which has been found to protect against mortality in patients with severe COVID-19, act by dampening this over-exuberant immune response ([RECOVERY trial, NEJM 2020](https://www.nejm.org/doi/full/10.1056/NEJMoa2021436)). Antibodies against the IL-6 receptor, such as tocilizumab and sarilumab were tested in clinical trials, but did not show efficacy (tocilizumab: [Stone et al., NEJM 2020](https://pubmed.ncbi.nlm.nih.gov/33085857/); sarilumab: [Sanofi press release 9/1/2020](https://www.sanofi.com/en/media-room/press-releases/2020/2020-09-01-07-00-00)), suggesting that a more narrow inhibition of IL-6 is not sufficient to treat COVID-19.
*Thought question:*
* How might the integrity of the lung and ability of immune cells to migrate to the site of infection affect the immune response to SARS-CoV-2?
### Adaptive immune response
The adaptive immune response generally consists of humoral immunity, most prominently antibodies produced by B cells, and cellular immunity, including CD4+ and CD8+ T cells and NK cells. These cells are primed by antigen presentation from cells including dendritic cells, macrophages, and B cells. One study using serological methods found low to no detection of SARS-CoV-2 specific antibodies in samples banked prior to SARS-CoV-2 exposure with negligible cross-reactivity from other human coronaviruses, suggesting humans may be completely immunologically naive to SARS-CoV-2 prior to the emergence of COVID-19 ([Amanat et al. Nature Medicine 2020](https://www.nature.com/articles/s41591-020-0913-5)). Once exposed, people can form a detectable humoral immune response to SARS-CoV-2. IgM antibodies to SARS-CoV-2 were found at a median of 12 days after illness onset and IgG after a median of 14 days; the timing of antibody production was not associated with disease severity, but higher antibody titer was associated with worse disease ([Zhao et al., Clinical Infectious Diseases 2020](https://academic.oup.com/cid/advance-article/doi/10.1093/cid/ciaa344/5812996)). Antibody responses are often of low magnitude ([Robbiani et al., 2020](https://www.nature.com/articles/s41586-020-2456-9)) and appear to lack durability ([Long et al., 2020](https://www.nature.com/articles/s41591-020-0897-1)). Antibody somatic hypermutation levels are low in both mild and severe disease, and autopsy studies have revealed the absence of germinal centers in patients with severe disease ([Kaneko et al. Cell 2020](https://www.cell.com/cell/pdf/S0092-8674\(20\)31067-9.pdf)). These results may explain the lack of durability of antibody responses in severely ill patients. The presence of an antibody and T cell response that protects from reinfection has been established in animal models ([Chandrashekar et al., Science, 2020](https://science.sciencemag.org/content/369/6505/812)). Data from a COVID-19 outbreak on a fishing boat found that the presence of neutralizing antibodies from a prior infection was protective against reinfection ([Addetia et al., J Clin Micro, 2020](https://jcm.asm.org/content/58/11/e02107-20)).
Neutralizing monoclonal antibodies to SARS-CoV2 have been cloned from humanized mice and humans ([Hansen et al., Science 2020](https://science.sciencemag.org/content/369/6506/1010?ijkey=f5391435661ccffed6e3690ce2ec1b66c796f753\&keytype2=tf_ipsecsha); [Baum et al., Science 2020](https://science.sciencemag.org/content/369/6506/1014)). Monoclonal antibodies from Lilly and Regeneron have both received emergency use authorization from the FDA for outpatient use ([Lilly press release 11/9/20](https://www.lilly.com/news/media/media-kits/bamlanivimab-covid19); [Regeneron press release 11/21/20](https://newsroom.regeneron.com/news-releases/news-release-details/regenerons-regen-cov2-first-antibody-cocktail-covid-19-receive)). . The Lilly antibody, called bamlanivimab, consists of one monoclonal antibody, while the Regeneron cocktail, called casirivimab and imdevimab, has two antibodies which bind to separate sites on the Spike protein, a combination which prevented development of resistance in vitro ([Baum et al., Science 2020).](https://science.sciencemag.org/content/369/6506/1014) Both companies’ monoclonal antibodies showed preliminary efficacy in decreasing viral loads and follow-up visits in outpatient clinical trials ([Regeneron press release 10/28/20](https://investor.regeneron.com/news-releases/news-release-details/regenerons-covid-19-outpatient-trial-prospectively-demonstrates); [Chen et al. for the BLAZE-1 study, NEJM 2020](https://www.nejm.org/doi/full/10.1056/NEJMoa2029849)). Lilly’s antibody, bamlanivimab, received an emergency use authorization for outpatient use from the FDA (). A parallel trial of bamlanivimab in hospitalized patients was halted for lack of efficacy ([New York Times 10/26/20](https://www.nytimes.com/live/2020/10/26/world/covid-19-coronavirus-updates/eli-lilly-said-its-antibody-treatment-does-not-work-on-patients-hospitalized-with-covid-19)), which suggests that the timing of antibody administration is important in efficacy. Immunologically, antiviral antibodies are most likely to be effective during earlier stages of infection where viral load is higher, rather than after critical illness sets in. However, it may be challenging to identify patients who would benefit most from these monoclonal antibodies, and the infusions are expensive and may be challenging to administer in the outpatient setting while maintaining infection precautions ([Sax, NEJM Journal Watch, 11/15/20](https://blogs.jwatch.org/hiv-id-observations/index.php/bamlanivimab-hard-to-pronounce-even-harder-to-give/2020/11/15/)).Convalescent plasma is another strategy for transferring protective antibodies to people with COVID-19. The FDA granted an emergency use authorization for convalescent plasma based on retrospective, indirect analyses of an open label protocol from the Mayo Clinic, which showed lower 7-day mortality in non-intubated patients who received high-titer plasma (11%) compared to low-titer plasma (14%) ([FDA EUA request](https://www.fda.gov/media/141481/download)). However, many other studies had equivocal results, and the NIH COVID-19 Treatment Guidelines currently state that there is insufficient data to recommend for or against use of convalescent plasma ([NIH COVID-19 Treatment Guidelines](https://www.covid19treatmentguidelines.nih.gov/immune-based-therapy/blood-derived-products/convalescent-plasma/)).
When it comes to T cells, CD4+ T cell responses against SARS-CoV-2 peptides are seen in 100% of COVID-19 patients, and CD8+ T cell responses are seen in 70% of these patients ([Grifoni et al., Cell 2020](https://www.cell.com/cell/pdf/S0092-8674\(20\)30610-3.pdf)). Interestingly, CD4+ T cells that react against SARS-CoV-2 were also seen in 40-60% of patient cells collected in 2015-2018, which correlated with the presence of antibodies against seasonal coronaviruses, suggesting pre-existing cross-immunity from other coronavirus infections ([Grifoni et al., Cell 2020](https://www.cell.com/cell/pdf/S0092-8674\(20\)30610-3.pdf)). However differentiation of T cells into Bcl-6+ T follicular helper cells is defective in severe COVID-19, and this likely compromises the quality of the humoral immune response ([Kaneko et al. Cell 2020](https://www.cell.com/cell/pdf/S0092-8674\(20\)31067-9.pdf)). A Th1-type CD4+ T cell response is important in successful control of SARS-CoV and MERS-CoV ([Li et al., J Imm 2008](https://www.jimmunol.org/content/jimmunol/181/8/5490.full.pdf); [Shin et al., Clin Inf Dis 2019](https://academic.oup.com/cid/article/68/6/984/5060259)). In mouse models of SARS, loss of CD4+ T cells reduced viral clearance, decreased antibody responses and led to increased mortality, while depletion of CD8+ T cells had no effect ([Shen et al., J Virol 2009](https://jvi.asm.org/content/84/3/1289.short)). In mice, CD4+ T resident memory cells in the lung are particularly important for vaccine-mediated protection against SARS ([Zhao et al., Immunity 2016](https://www.sciencedirect.com/science/article/pii/S1074761316301601)). CD8+ T cell responses are also important to control infection, but may be associated with increased lung pathology in SARS and MERS when overabundant, making it difficult to discern cause from consequence ([Shin et al., Clin Inf Dis 2019](https://academic.oup.com/cid/article/68/6/984/5060259); [Prompetchara et al., As Pac J of All and Imm 2020](http://apjai-journal.org/wp-content/uploads/2020/02/Covid_AP-200220-0772.pdf)). MERS-CoV has been shown to decrease antigen presentation on dendritic cells and macrophages, delaying activation of the adaptive immune system ([Shokri et al., J Cell Physiol, 2019](https://onlinelibrary.wiley.com/doi/full/10.1002/jcp.27155)).
Lymphopenia is a hallmark of in COVID-19, is associated with IL-6 and IL-8 levels ([Zhang et al., Nature 2020](https://www.nature.com/articles/s41586-020-2355-0)), and is a predictor of disease severity ([Tan et al., Signal Transduction and Targeted Therapy 2020](https://www.nature.com/articles/s41392-020-0148-4#MOESM1)). CD4+ and CD8+ T cells are depleted to a greater extent than B cells or NK cells ([Zhang et al., Nature 2020](https://www.nature.com/articles/s41586-020-2355-0)), which may be partially explained by recruitment to infected tissue. This may be due to bone marrow suppression by the antiviral response, destruction of lymphatic tissue ([Chen et al., MedRXiv 2020](https://www.medrxiv.org/content/10.1101/2020.03.27.20045427v1)), or perhaps by direct viral infection and depletion of lymphocytes ([Wang et al., Cellular and Mol Immunol, 2020](https://www.nature.com/articles/s41423-020-0424-9)). It is not yet known whether individuals with mild disease recover in part because of vigorous protective cytotoxic CD8+ T cell based elimination of infected cells, and whether the progression of illness in more seriously ill patients reflects a relative absence of such a response. These differences, if established, could in part help explain the clinical spectrum of the disease.\
\
A key question for understanding the dynamics of the pandemic is whether previously infected individuals can be re-infected. It appears that re-infection is possible but rare. In macaque models of infection, some monkeys become infected again after rechallenge, but this infection only lasted a short period of time and no infectious virus was recovered, suggesting that they had developed an effective immune response ([Chandrashekar et al., Science 2020](https://science.sciencemag.org/content/early/2020/05/19/science.abc4776.abstract)). A case of human reinfection 142 days after the initial infection with a genetically distinct viral strain has been reported; however, the patient was asymptomatic during the second episode ([To et al., Clin Inf Dis 2020](https://academic.oup.com/cid/advance-article/doi/10.1093/cid/ciaa1275/5897019)). Cases of human reinfection (people testing positive for SARS-CoV-2 RNA after testing negative) are most likely rare; a study of 790 contacts of 285 re-positive cases in South Korea found no cases of transmission from re-positive individuals, so they are very unlikely to be infectious ([Korean CDC, 5/21/20](https://www.cdc.go.kr/board/board.es?mid=a30402000000\&bid=0030\&act=view\&list_no=367267\&nPage=1)). Immunity to seasonal coronaviruses wanes over several years, permitting reinfection with very similar strains of the same virus, and antibody titers to SARS and MERS decrease after infection, raising questions about the durability of protection against COVID-19 . Encouragingly, a study that sampled immune responses to SARS-CoV-2 in people with mild, moderate and severe COVID-19 over time with up to 8 months of follow-up found persistence of antibody, B cell and T cell responses ([Dan et al., BioRXiv 11/16/20](https://www.biorxiv.org/content/10.1101/2020.11.15.383323v1.full.pdf)). Data from phase 3 clinical trials from currently approved vaccine candidates show that antibody and T cell responses are durable and that protection can last for at least several months. The long-term duration of protection from previous infection and vaccines will become more apparent as time goes on.\
\
For more detailed information on vaccine development, see the below [section](https://docs.google.com/document/d/1wXMH05bfCrj4UFXtJ_DFZE9g_WrEjj7V7MB_Ur7l7m0/edit#bookmark=kix.th8bz2lrbutu). For more information about how this affects epidemiological modeling, see [Module 2](/module-2-epidemiology-principles.md#introduction).
*Thought question:*
* How might the initial mild presentation and later severe disease seen in COVID-19 be explained by the immune response to the virus?
Additional Readings:
[Prompetchara et al., Asian Pacific Journal of Allergy and Immunology, 2020](http://apjai-journal.org/wp-content/uploads/2020/02/Covid_AP-200220-0772.pdf)\
[de Wit et al., Nat Rev Microbiol 2016](https://www.nature.com/articles/nrmicro.2016.81.pdf)
---
# Agent Instructions: Querying This Documentation
If you need additional information that is not directly available in this page, you can query the documentation dynamically by asking a question.
Perform an HTTP GET request on the current page URL with the `ask` query parameter:
```
GET https://curriculum.covidstudentresponse.org/module-1-from-bench-to-bedside/basic-virology-and-immunology.md?ask=
```
The question should be specific, self-contained, and written in natural language.
The response will contain a direct answer to the question and relevant excerpts and sources from the documentation.
Use this mechanism when the answer is not explicitly present in the current page, you need clarification or additional context, or you want to retrieve related documentation sections.